The peptide bond thus has significant double-bond character (bond length 1.32 Å) which inhibits rotation about the N– CO bond: in this respect amides resemble alkenes. A key structural feature of polypeptides and proteins is the planarity of an amide bond, in which the N-atom has sp 2 (trigonal planar) geometry. The geometry is strongly influenced by two factors: The term secondary structure refers to the geometry of a polypeptide's N-C-C backbone, which can be randomly coiled or may be ordered in special ways. The nature of these is discussed in the following sections. Proteins are not only long chains, but fold up into complex 3-D structures that include lengths of randomly coiled chain along with regions of ordered structure. Nature clearly has many options for the construction of proteins from different permutations of the 20-odd amino acids that are routinely available.īecause the α-amino acids are single enantiomers, the polypeptides and proteins that are formed from them are also chiral (handed) molecules.
An octapeptide has 40,320 possible primary structures. It is this sequence of amino acids in a polypeptide chain that is referred to as its primary structure. In general, when polymerising n different amino acids (using each once) there are n! possible sequences of product.
For example, the pentapeptide met-enkephalin can be written as H-Tyr-Gly-Gly-Phe-Met-OH. Three-letter codes are very useful for describing polypeptides without having to draw them. The amino acid at the left-hand end is referred to as the N -terminal amino acid, and the amino acid at the right-hand end is referred to as the C -terminal amino acid. The convention for drawing peptides is to put the N-terminus on the left. When the new amide bond involves two of the natural amino acids, it is referred to as a peptide bond. These can form amides by the condensation of the amine group of one amino acid with the carboxylic acid group of another. The building blocks of proteins are approximately twenty naturally occurring alpha-amino acids (for structures, see Appendix 3: Proteinogenic Amino Acids). One distinction is between simple and conjugated proteins: Simple proteins are composed entirely of amino acids conjugated proteins have a non-amino acid component (called a prosthetic group). Proteins can be classified with reference to any of several distinguishing features (molecular weight, occurrence, solubility, etc.). Some useful definitions of these roles are shown in the box on the right.
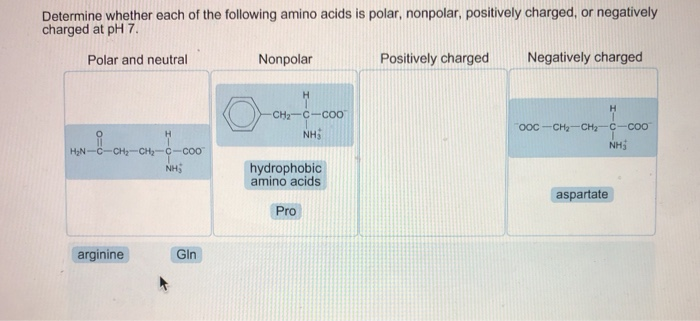
ARE HYDROPHOBIC AMINO ACIDS POLAR OR NONPOLAR CODE
Thus, a simple binary code of polar and nonpolar residues arranged in the appropriate order can drive polypeptide chains to collapse into globular α-helical folds.Proteins are polymeric amino acids of high molecular weight, arbitrarily 10,000 Da or more, which fulfil important roles in an organism's life cycle. Characterization of the expressed proteins indicated that most of the designed sequences folded into compact α-helical structures. Each gene encoded a different amino acid sequence, but all sequences shared the same pattern of polar and nonpolar residues. This strategy was tested by constructing a large collection of synthetic genes whose protein products were designed to fold into four-helix bundle proteins. In this strategy the sequence locations of hydrophobic and hydrophilic residues were specified explicitly, but the precise identities of the side chains were not constrained and varied extensively. Thus, a simple binary code of polar and nonpolar residues arranged in the appropriate order can drive polypeptide chains to collapse into globular α-helical folds.ĪB - A general strategy is described for the de novo design of proteins. N2 - A general strategy is described for the de novo design of proteins. T1 - Protein design by binary patterning of polar and nonpolar amino acids
